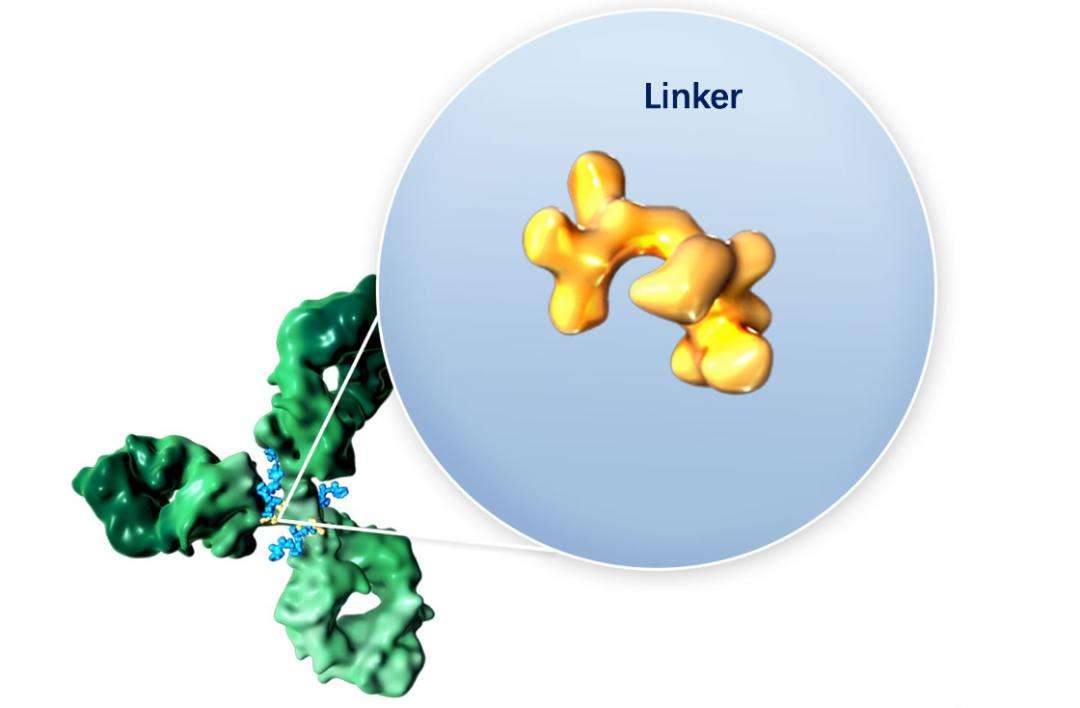
1. Conjugation of Lysine Residues
This method attaches the payload to the lysine residue of the antibody through a linker containing an active carboxylate site.
2. Reduction of cysteine coupling by disulfide bond modification
Taking IgG1 as an example, there are 4 pairs of interchain disulfide bonds that can be reduced, and 8 cysteine sulfhydryl groups are obtained after reduction. Groups such as maleimide on Linker can react with sulfhydryl groups to form stable conjugates. The DAR value of the final product can be optimized by the degree of disulfide reduction
3. Unnatural amino acid conjugation
This method introduces the unnatural amino acid into the antibody by constructing an unnatural amino acid expression system (for example, using tRNA carrying the unnatural amino acid, etc.), and connects with the linker at the unnatural amino acid site.
4. Enzyme-catalyzed coupling
In this method, the specific amino acid sequence on the antibody is recognized by the enzyme, and the corresponding site is modified to generate a conjugation site.
5. Coupling via transpeptidase-mediated transpeptidation
This method relies on the catalytic properties of the transpeptidase. Using this feature of Sortase A, various types of molecules can be attached to oligo-G to achieve conjugation to specific sites of antibodies.
6. Coupling via MTGase (microbial transglutaminase)-mediated transpeptidation
The primary amine-containing linker is covalently attached to the primary amine of the specific glutamine (Q295) of the deglycosylated antibody using an MTGase-catalyzed transpeptidation reaction, since each heavy chain of the antibody has one binding site, Therefore, the DAR value of ADC molecules obtained by this method is fixed at 2.
7. Conjugation by engineering the N-glycan on the aspartic acid residue of the antibody.